Research team develops new tools to improve pancreatic cancer patient care

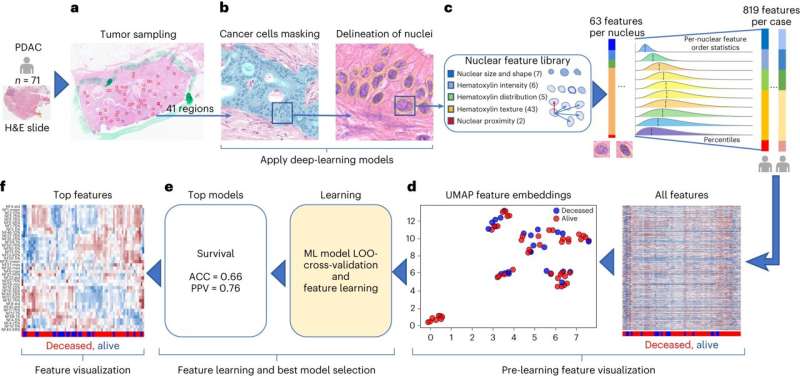
Cedars-Sinai Cancer investigators have used a unique precision medicine and artificial intelligence (AI) tool called the Molecular Twin Precision Oncology Platform to identify biomarkers that outperform the standard test for predicting pancreatic cancer survival. Their study, published in Nature Cancer, demonstrates the viability of a tool that could one day guide and improve treatment for all cancer patients.
“Molecular Twin, which we developed at Cedars-Sinai, can be used to study any tumor type, including pancreatic cancer, which is notoriously difficult to treat,” said Dan Theodorescu, MD, Ph.D., director of Cedars-Sinai Cancer and the PHASE ONE Foundation Distinguished Chair, and senior author of the study. “Using our Molecular Twin technology, we anticipate creating tests that can be used even in locations that lack access to advanced resources and technology, pairing patients with the most effective therapies and expanding the availability of precision medicine.”
Investigators used the Molecular Twin platform to analyze blood and tissue samples from 74 patients with the most common and most aggressive pancreatic cancer type, pancreatic ductal adenocarcinoma. The disease begins in the cells lining ducts that carry digestive enzymes from the pancreas to the small intestine.
Investigators first combined 6,363 different biological data points, including genetic and molecular information, to create a model that accurately predicted disease survival in 87% of patients. The team then used AI to streamline the data and create a model that performed nearly as well with just 589 points of data. Zeroing in even further, investigators determined that proteins found in the blood were the best single predictor of pancreatic cancer survival.
The full and streamlined models, and the blood-protein test, outperformed the only Food and Drug Administration-approved pancreatic cancer test, a blood test called CA 19-9. The findings were validated in independent datasets from The Cancer Genome Atlas, Massachusetts General Hospital and Johns Hopkins University.
The Molecular Twin platform was launched by Cedars-Sinai in 2021, said Arsen Osipov, MD, assistant professor of Medicine, program lead in the Pancreatic Cancer Multidisciplinary Clinic and Precision Medicine Program at Cedars-Sinai Cancer, and first author of the study.
“There’s a huge unmet need for the development of biomarkers to guide our treatment of pancreatic cancer,” Osipov said. “We had already undertaken a comprehensive collection of blood and tissue samples from patients with pancreatic cancer, and this gave us a good opportunity to test the Molecular Twin platform. As we grow the platform with more patients, Molecular Twin will become an even more robust tool, not just in pancreatic cancer, but across all cancers.”
Jennifer Van Eyk, Ph.D., an expert in the study of proteins, director of the Advanced Clinical Biosystems Institute in the Department of Biomedical Sciences at Cedars-Sinai and a key member of the Molecular Twin team, said that while genetic information is helpful in predicting a patient’s risk of developing cancer and the subtyping of the cancer, this study shows that proteins are the key to predicting patient survival.
“Once a patient has cancer, proteins act as the body’s first responders, and their activity helps us determine how a patient’s body is reacting,” Van Eyk said. “Proteins turned out to be the main drivers of our pancreatic cancer models. And in future studies, proteins will also help us track how well a patient is responding to treatment.”
While this initial use of Molecular Twin aims to develop tests that guide treatment of pancreatic cancer patients, the platform and its uses will continue to expand, Theodorescu said. Investigators are adding data from additional patients and branching out to include additional types of data such as medical imaging, samples of the gut microbiome and tumor microenvironment, and feedback from wearable devices that measure physical activity.
“A majority of our cancer patients are allowing us to include their clinical information and samples from blood, tumor and other sources so that we can continue to build the Molecular Twin platform,” Theodorescu said. “This rich pool of data will help us discover biomarkers for additional cancer types, and eventually lead to the development of new treatments and the opportunity to identify at-risk patients before their cancer develops, so that we can prevent it entirely.”
More information:
Arsen Osipov et al, The Molecular Twin artificial-intelligence platform integrates multi-omic data to predict outcomes for pancreatic adenocarcinoma patients, Nature Cancer (2024). DOI: 10.1038/s43018-023-00697-7
Cedars-Sinai Medical Center
Citation:
Research team develops new tools to improve pancreatic cancer patient care (2024, January 22)
retrieved 29 January 2024
from
This document is subject to copyright. Apart from any fair dealing for the purpose of private study or research, no
part may be reproduced without the written permission. The content is provided for information purposes only.
link